Chapter 13 Novelty-lexicase
13.1 Overview
13.2 Analysis dependencies
library(ggplot2)
library(tidyverse)
library(knitr)
library(cowplot)
library(viridis)
library(RColorBrewer)
library(rstatix)
library(ggsignif)
library(Hmisc)
library(kableExtra)
source("https://gist.githubusercontent.com/benmarwick/2a1bb0133ff568cbe28d/raw/fb53bd97121f7f9ce947837ef1a4c65a73bffb3f/geom_flat_violin.R")These analyses were conducted in the following computing environment:
## _
## platform x86_64-pc-linux-gnu
## arch x86_64
## os linux-gnu
## system x86_64, linux-gnu
## status
## major 4
## minor 1.0
## year 2021
## month 05
## day 18
## svn rev 80317
## language R
## version.string R version 4.1.0 (2021-05-18)
## nickname Camp Pontanezen13.3 Setup
data_loc <- paste0(
working_directory,
"data/timeseries-res-1000g.csv"
)
data <- read.csv(data_loc, na.strings="NONE")
data$cardinality <- as.factor(
data$OBJECTIVE_CNT
)
data$selection_name <- as.factor(
data$selection_name
)
data$epsilon <- as.factor(
data$LEX_EPS
)
data$k <- as.factor(
data$NOVEL_K
)
data$elite_trait_avg <-
data$ele_agg_per / data$OBJECTIVE_CNT
data$unique_start_positions_coverage <-
data$uni_str_pos / data$OBJECTIVE_CNT
final_data <- filter(data, evaluations==max(data$evaluations))
# Labeler for stats annotations
p_label <- function(p_value) {
threshold = 0.0001
if (p_value < threshold) {
return(paste0("p < ", threshold))
} else {
return(paste0("p = ", p_value))
}
}
# Significance threshold
alpha <- 0.05
####### misc #######
# Configure our default graphing theme
theme_set(theme_cowplot())13.4 Exploration diagnostic performance
elite_ave_performance_fig <-
ggplot(
data,
aes(x=gen, y=elite_trait_avg, color=k, fill=k)
) +
stat_summary(geom="line", fun=mean) +
stat_summary(
geom="ribbon",
fun.data="mean_cl_boot",
fun.args=list(conf.int=0.95),
alpha=0.2,
linetype=0
) +
scale_y_continuous(
name="Average trait performance",
limits=c(0, 100)
) +
scale_x_continuous(
name="Generations"
) +
scale_fill_brewer(
name="k",
palette=cb_palette
) +
scale_color_brewer(
name="k",
palette=cb_palette
)
elite_ave_performance_fig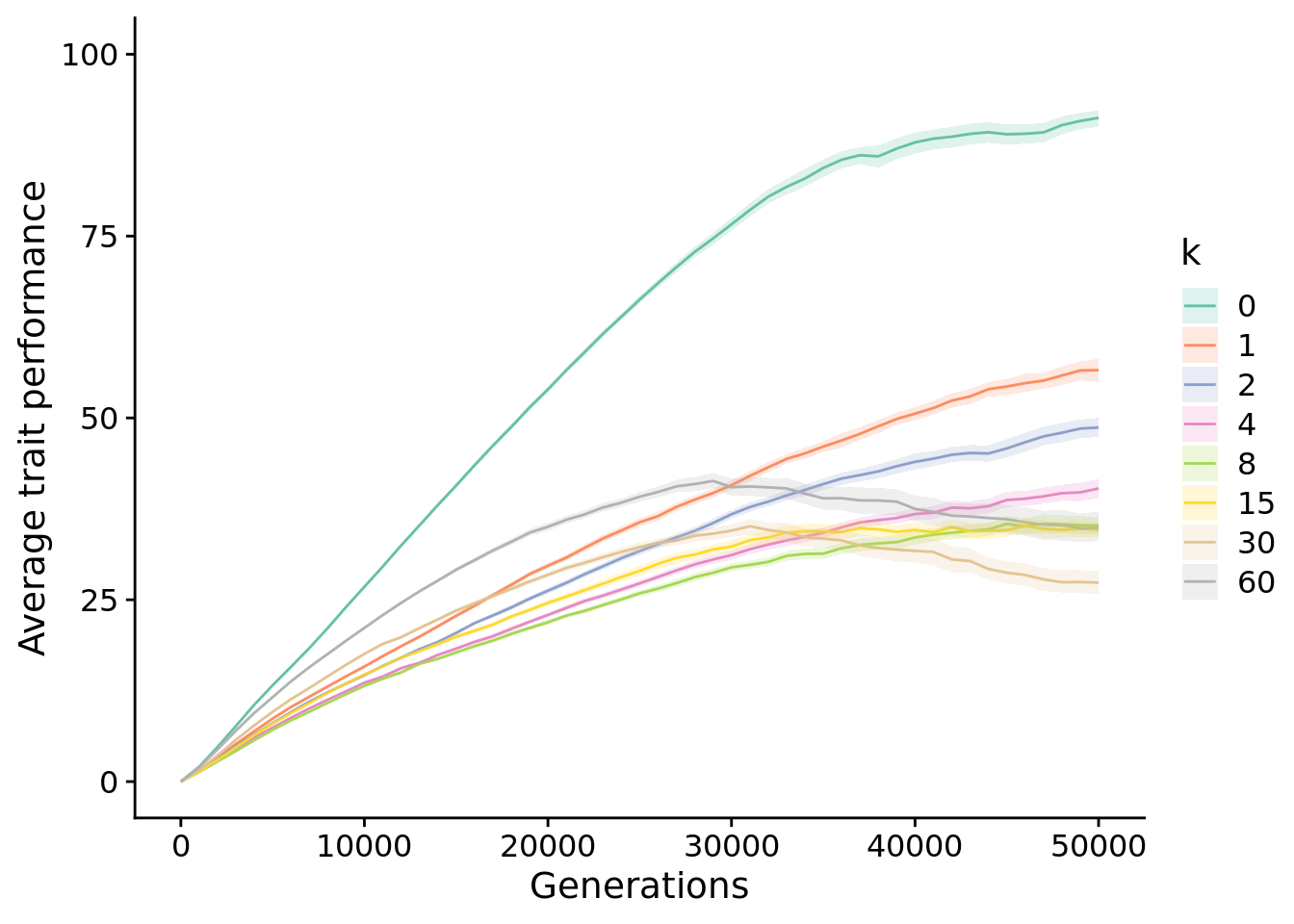
13.4.1 Final performance
# Compute manual labels for geom_signif
stat.test <- final_data %>%
wilcox_test(elite_trait_avg ~ k) %>%
adjust_pvalue(method = "bonferroni") %>%
add_significance() %>%
add_xy_position(x="k",step.increase=1)
stat.test$manual_position <- stat.test$y.position * 1.05
stat.test$label <- mapply(p_label,stat.test$p.adj)elite_final_performance_fig <- ggplot(
final_data,
aes(x=k, y=elite_trait_avg, fill=k)
) +
geom_flat_violin(
position = position_nudge(x = .2, y = 0),
alpha = .8,
scale="width"
) +
geom_point(
mapping=aes(color=k),
position = position_jitter(width = .15),
size = .5,
alpha = 0.8
) +
geom_boxplot(
width = .1,
outlier.shape = NA,
alpha = 0.5
) +
scale_y_continuous(
name="Average trait performance",
limits=c(0, 100)
) +
scale_x_discrete(
name="k"
) +
scale_fill_brewer(
name="k",
palette=cb_palette
) +
scale_color_brewer(
name="k",
palette=cb_palette
) +
theme(
legend.position="none"
)
elite_final_performance_fig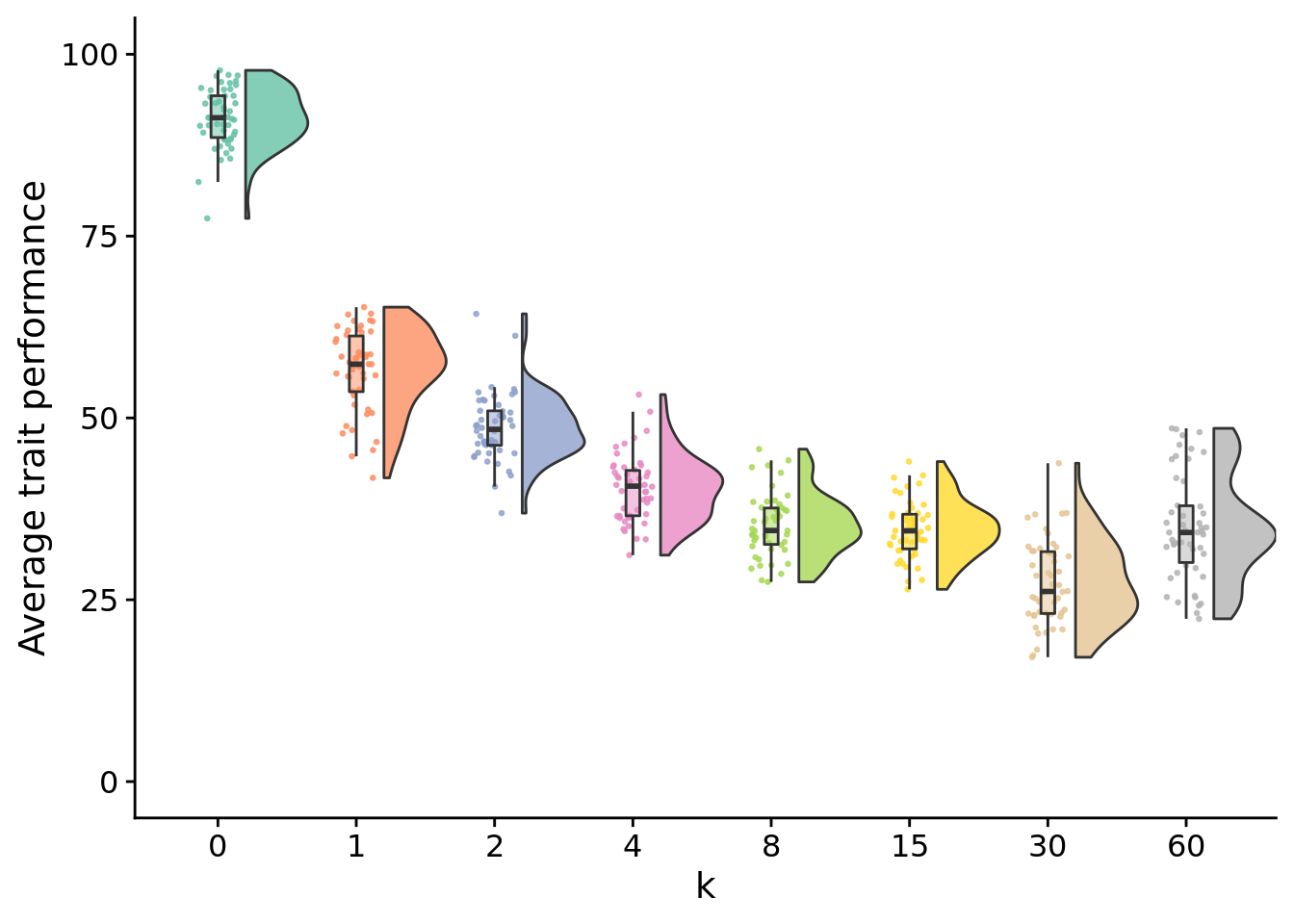
stat.test %>%
kbl() %>%
kable_styling(
bootstrap_options = c(
"striped",
"hover",
"condensed",
"responsive"
)
) %>%
scroll_box(width="600px")| .y. | group1 | group2 | n1 | n2 | statistic | p | p.adj | p.adj.signif | y.position | groups | xmin | xmax | manual_position | label |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| elite_trait_avg | 0 | 1 | 50 | 50 | 2500 | 0.00e+00 | 0.0000000 | **** | 151.8030 | 0, 1 | 1 | 2 | 159.3931 | p < 1e-04 |
| elite_trait_avg | 0 | 2 | 50 | 50 | 2500 | 0.00e+00 | 0.0000000 | **** | 207.8186 | 0, 2 | 1 | 3 | 218.2095 | p < 1e-04 |
| elite_trait_avg | 0 | 4 | 50 | 50 | 2500 | 0.00e+00 | 0.0000000 | **** | 263.8341 | 0, 4 | 1 | 4 | 277.0258 | p < 1e-04 |
| elite_trait_avg | 0 | 8 | 50 | 50 | 2500 | 0.00e+00 | 0.0000000 | **** | 319.8497 | 0, 8 | 1 | 5 | 335.8422 | p < 1e-04 |
| elite_trait_avg | 0 | 15 | 50 | 50 | 2500 | 0.00e+00 | 0.0000000 | **** | 375.8652 | 0 , 15 | 1 | 6 | 394.6585 | p < 1e-04 |
| elite_trait_avg | 0 | 30 | 50 | 50 | 2500 | 0.00e+00 | 0.0000000 | **** | 431.8808 | 0 , 30 | 1 | 7 | 453.4748 | p < 1e-04 |
| elite_trait_avg | 0 | 60 | 50 | 50 | 2500 | 0.00e+00 | 0.0000000 | **** | 487.8963 | 0 , 60 | 1 | 8 | 512.2912 | p < 1e-04 |
| elite_trait_avg | 1 | 2 | 50 | 50 | 2137 | 0.00e+00 | 0.0000000 | **** | 543.9119 | 1, 2 | 2 | 3 | 571.1075 | p < 1e-04 |
| elite_trait_avg | 1 | 4 | 50 | 50 | 2451 | 0.00e+00 | 0.0000000 | **** | 599.9274 | 1, 4 | 2 | 4 | 629.9238 | p < 1e-04 |
| elite_trait_avg | 1 | 8 | 50 | 50 | 2493 | 0.00e+00 | 0.0000000 | **** | 655.9430 | 1, 8 | 2 | 5 | 688.7401 | p < 1e-04 |
| elite_trait_avg | 1 | 15 | 50 | 50 | 2497 | 0.00e+00 | 0.0000000 | **** | 711.9586 | 1 , 15 | 2 | 6 | 747.5565 | p < 1e-04 |
| elite_trait_avg | 1 | 30 | 50 | 50 | 2499 | 0.00e+00 | 0.0000000 | **** | 767.9741 | 1 , 30 | 2 | 7 | 806.3728 | p < 1e-04 |
| elite_trait_avg | 1 | 60 | 50 | 50 | 2467 | 0.00e+00 | 0.0000000 | **** | 823.9897 | 1 , 60 | 2 | 8 | 865.1892 | p < 1e-04 |
| elite_trait_avg | 2 | 4 | 50 | 50 | 2277 | 0.00e+00 | 0.0000000 | **** | 880.0052 | 2, 4 | 3 | 4 | 924.0055 | p < 1e-04 |
| elite_trait_avg | 2 | 8 | 50 | 50 | 2459 | 0.00e+00 | 0.0000000 | **** | 936.0208 | 2, 8 | 3 | 5 | 982.8218 | p < 1e-04 |
| elite_trait_avg | 2 | 15 | 50 | 50 | 2480 | 0.00e+00 | 0.0000000 | **** | 992.0363 | 2 , 15 | 3 | 6 | 1041.6381 | p < 1e-04 |
| elite_trait_avg | 2 | 30 | 50 | 50 | 2494 | 0.00e+00 | 0.0000000 | **** | 1048.0519 | 2 , 30 | 3 | 7 | 1100.4545 | p < 1e-04 |
| elite_trait_avg | 2 | 60 | 50 | 50 | 2341 | 0.00e+00 | 0.0000000 | **** | 1104.0674 | 2 , 60 | 3 | 8 | 1159.2708 | p < 1e-04 |
| elite_trait_avg | 4 | 8 | 50 | 50 | 1988 | 4.00e-07 | 0.0000103 | **** | 1160.0830 | 4, 8 | 4 | 5 | 1218.0872 | p < 1e-04 |
| elite_trait_avg | 4 | 15 | 50 | 50 | 2074 | 0.00e+00 | 0.0000004 | **** | 1216.0986 | 4 , 15 | 4 | 6 | 1276.9035 | p < 1e-04 |
| elite_trait_avg | 4 | 30 | 50 | 50 | 2389 | 0.00e+00 | 0.0000000 | **** | 1272.1141 | 4 , 30 | 4 | 7 | 1335.7198 | p < 1e-04 |
| elite_trait_avg | 4 | 60 | 50 | 50 | 1854 | 3.18e-05 | 0.0008904 | *** | 1328.1297 | 4 , 60 | 4 | 8 | 1394.5361 | p = 0.0008904 |
| elite_trait_avg | 8 | 15 | 50 | 50 | 1343 | 5.24e-01 | 1.0000000 | ns | 1384.1452 | 8 , 15 | 5 | 6 | 1453.3525 | p = 1 |
| elite_trait_avg | 8 | 30 | 50 | 50 | 2159 | 0.00e+00 | 0.0000000 | **** | 1440.1608 | 8 , 30 | 5 | 7 | 1512.1688 | p < 1e-04 |
| elite_trait_avg | 8 | 60 | 50 | 50 | 1346 | 5.10e-01 | 1.0000000 | ns | 1496.1763 | 8 , 60 | 5 | 8 | 1570.9851 | p = 1 |
| elite_trait_avg | 15 | 30 | 50 | 50 | 2127 | 0.00e+00 | 0.0000000 | **** | 1552.1919 | 15, 30 | 6 | 7 | 1629.8015 | p < 1e-04 |
| elite_trait_avg | 15 | 60 | 50 | 50 | 1287 | 8.01e-01 | 1.0000000 | ns | 1608.2074 | 15, 60 | 6 | 8 | 1688.6178 | p = 1 |
| elite_trait_avg | 30 | 60 | 50 | 50 | 515 | 4.00e-07 | 0.0000115 | **** | 1664.2230 | 30, 60 | 7 | 8 | 1747.4342 | p < 1e-04 |
13.5 Activation position coverage
unique_start_position_coverage_fig <- ggplot(
data,
aes(x=gen, y=unique_start_positions_coverage, color=k, fill=k)
) +
stat_summary(geom="line", fun=mean) +
stat_summary(
geom="ribbon",
fun.data="mean_cl_boot",
fun.args=list(conf.int=0.95),
alpha=0.2,
linetype=0
) +
scale_y_continuous(
name="Activation position coverage",
limits=c(0.0, 1.0)
) +
scale_x_continuous(
name="Generations"
) +
scale_fill_brewer(
name="k",
palette=cb_palette
) +
scale_color_brewer(
name="k",
palette=cb_palette
)
unique_start_position_coverage_fig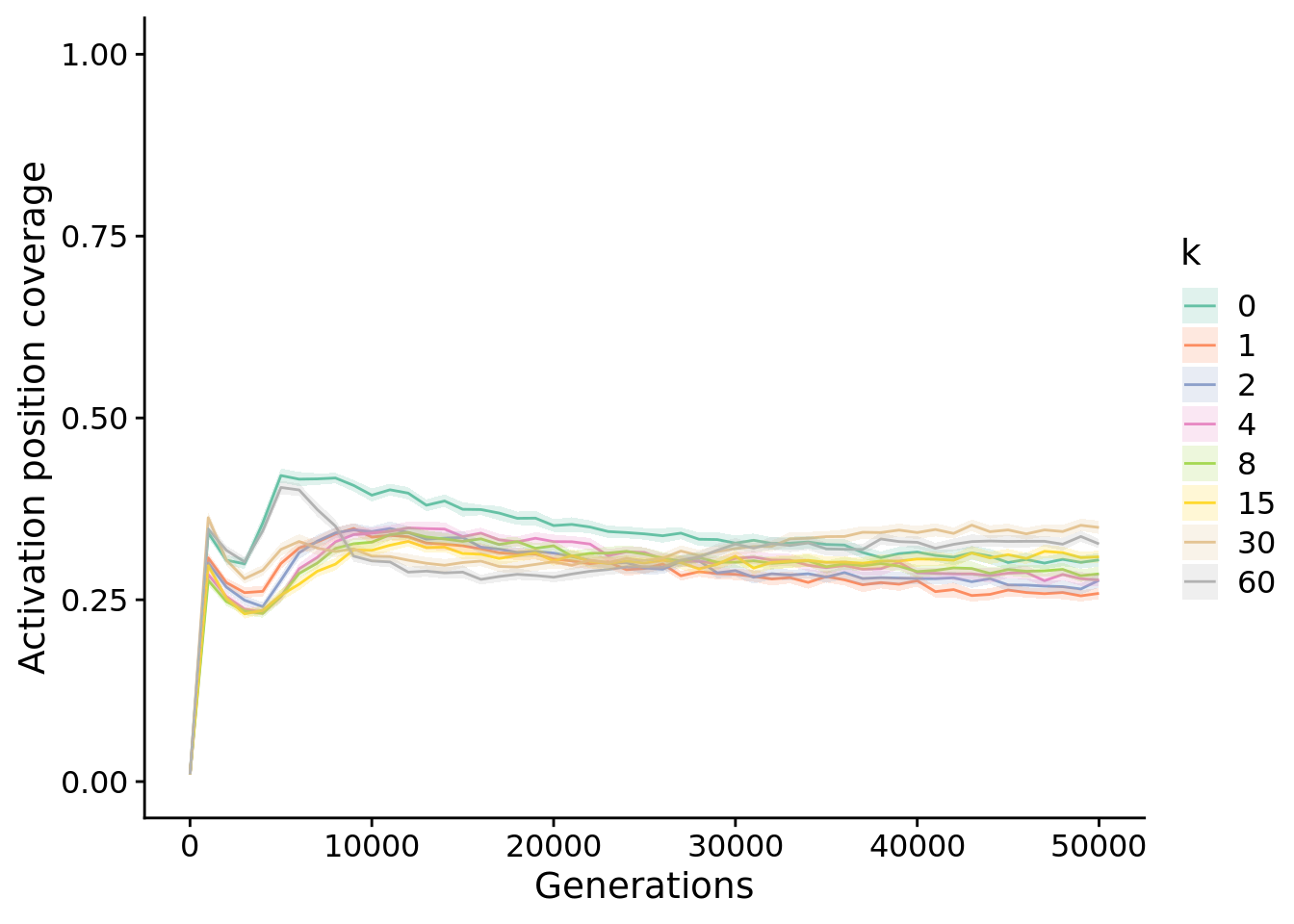
13.5.1 Final activation position coverage
# Compute manual labels for geom_signif
stat.test <- final_data %>%
wilcox_test(unique_start_positions_coverage ~ k) %>%
adjust_pvalue(method = "bonferroni") %>%
add_significance() %>%
add_xy_position(x="k",step.increase=1)
stat.test$manual_position <- stat.test$y.position * 1.05
stat.test$label <- mapply(p_label,stat.test$p.adj)unique_start_positions_coverage_final_fig <- ggplot(
final_data,
aes(x=k, y=unique_start_positions_coverage, fill=k)
) +
geom_flat_violin(
position = position_nudge(x = .2, y = 0),
alpha = .8,
scale="width"
) +
geom_point(
mapping=aes(color=k),
position = position_jitter(width = .15),
size = .5,
alpha = 0.8
) +
geom_boxplot(
width = .1,
outlier.shape = NA,
alpha = 0.5
) +
scale_y_continuous(
name="Activation position coverage",
limits=c(0, 1.0)
) +
scale_x_discrete(
name="k"
) +
scale_fill_brewer(
name="k",
palette=cb_palette
) +
scale_color_brewer(
name="k",
palette=cb_palette
) +
theme(
legend.position="none"
)
unique_start_positions_coverage_final_fig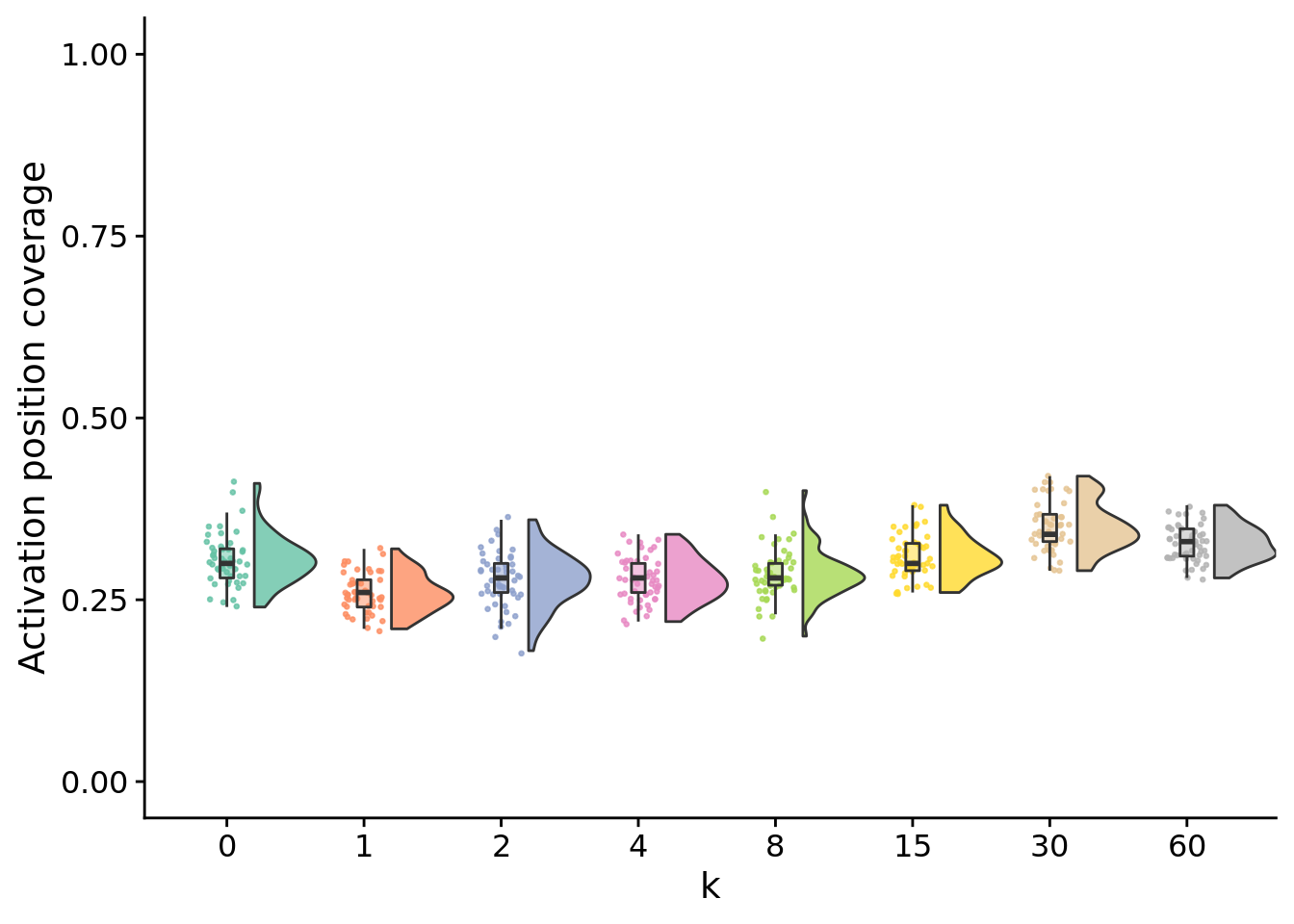
stat.test %>%
kbl() %>%
kable_styling(
bootstrap_options = c(
"striped",
"hover",
"condensed",
"responsive"
)
) %>%
scroll_box(width="600px")| .y. | group1 | group2 | n1 | n2 | statistic | p | p.adj | p.adj.signif | y.position | groups | xmin | xmax | manual_position | label |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| unique_start_positions_coverage | 0 | 1 | 50 | 50 | 2151.0 | 0.00e+00 | 0.0000000 | **** | 0.5200000 | 0, 1 | 1 | 2 | 0.5460000 | p < 1e-04 |
| unique_start_positions_coverage | 0 | 2 | 50 | 50 | 1759.5 | 4.26e-04 | 0.0119280 |
|
0.6237037 | 0, 2 | 1 | 3 | 0.6548889 | p = 0.011928 |
| unique_start_positions_coverage | 0 | 4 | 50 | 50 | 1806.0 | 1.19e-04 | 0.0033320 | ** | 0.7274074 | 0, 4 | 1 | 4 | 0.7637778 | p = 0.003332 |
| unique_start_positions_coverage | 0 | 8 | 50 | 50 | 1675.0 | 3.00e-03 | 0.0840000 | ns | 0.8311111 | 0, 8 | 1 | 5 | 0.8726667 | p = 0.084 |
| unique_start_positions_coverage | 0 | 15 | 50 | 50 | 1132.0 | 4.15e-01 | 1.0000000 | ns | 0.9348148 | 0 , 15 | 1 | 6 | 0.9815556 | p = 1 |
| unique_start_positions_coverage | 0 | 30 | 50 | 50 | 405.5 | 0.00e+00 | 0.0000001 | **** | 1.0385185 | 0 , 30 | 1 | 7 | 1.0904444 | p < 1e-04 |
| unique_start_positions_coverage | 0 | 60 | 50 | 50 | 692.5 | 1.13e-04 | 0.0031640 | ** | 1.1422222 | 0 , 60 | 1 | 8 | 1.1993333 | p = 0.003164 |
| unique_start_positions_coverage | 1 | 2 | 50 | 50 | 839.0 | 4.00e-03 | 0.1120000 | ns | 1.2459259 | 1, 2 | 2 | 3 | 1.3082222 | p = 0.112 |
| unique_start_positions_coverage | 1 | 4 | 50 | 50 | 820.0 | 3.00e-03 | 0.0840000 | ns | 1.3496296 | 1, 4 | 2 | 4 | 1.4171111 | p = 0.084 |
| unique_start_positions_coverage | 1 | 8 | 50 | 50 | 659.5 | 4.35e-05 | 0.0012180 | ** | 1.4533333 | 1, 8 | 2 | 5 | 1.5260000 | p = 0.001218 |
| unique_start_positions_coverage | 1 | 15 | 50 | 50 | 249.0 | 0.00e+00 | 0.0000000 | **** | 1.5570370 | 1 , 15 | 2 | 6 | 1.6348889 | p < 1e-04 |
| unique_start_positions_coverage | 1 | 30 | 50 | 50 | 33.0 | 0.00e+00 | 0.0000000 | **** | 1.6607407 | 1 , 30 | 2 | 7 | 1.7437778 | p < 1e-04 |
| unique_start_positions_coverage | 1 | 60 | 50 | 50 | 78.5 | 0.00e+00 | 0.0000000 | **** | 1.7644444 | 1 , 60 | 2 | 8 | 1.8526667 | p < 1e-04 |
| unique_start_positions_coverage | 2 | 4 | 50 | 50 | 1275.0 | 8.65e-01 | 1.0000000 | ns | 1.8681481 | 2, 4 | 3 | 4 | 1.9615556 | p = 1 |
| unique_start_positions_coverage | 2 | 8 | 50 | 50 | 1124.5 | 3.86e-01 | 1.0000000 | ns | 1.9718519 | 2, 8 | 3 | 5 | 2.0704444 | p = 1 |
| unique_start_positions_coverage | 2 | 15 | 50 | 50 | 631.0 | 1.84e-05 | 0.0005152 | *** | 2.0755556 | 2 , 15 | 3 | 6 | 2.1793333 | p = 0.0005152 |
| unique_start_positions_coverage | 2 | 30 | 50 | 50 | 169.0 | 0.00e+00 | 0.0000000 | **** | 2.1792593 | 2 , 30 | 3 | 7 | 2.2882222 | p < 1e-04 |
| unique_start_positions_coverage | 2 | 60 | 50 | 50 | 325.0 | 0.00e+00 | 0.0000000 | **** | 2.2829630 | 2 , 60 | 3 | 8 | 2.3971111 | p < 1e-04 |
| unique_start_positions_coverage | 4 | 8 | 50 | 50 | 1074.5 | 2.25e-01 | 1.0000000 | ns | 2.3866667 | 4, 8 | 4 | 5 | 2.5060000 | p = 1 |
| unique_start_positions_coverage | 4 | 15 | 50 | 50 | 576.5 | 3.10e-06 | 0.0000874 | **** | 2.4903704 | 4 , 15 | 4 | 6 | 2.6148889 | p < 1e-04 |
| unique_start_positions_coverage | 4 | 30 | 50 | 50 | 126.5 | 0.00e+00 | 0.0000000 | **** | 2.5940741 | 4 , 30 | 4 | 7 | 2.7237778 | p < 1e-04 |
| unique_start_positions_coverage | 4 | 60 | 50 | 50 | 270.0 | 0.00e+00 | 0.0000000 | **** | 2.6977778 | 4 , 60 | 4 | 8 | 2.8326667 | p < 1e-04 |
| unique_start_positions_coverage | 8 | 15 | 50 | 50 | 702.0 | 1.47e-04 | 0.0041160 | ** | 2.8014815 | 8 , 15 | 5 | 6 | 2.9415556 | p = 0.004116 |
| unique_start_positions_coverage | 8 | 30 | 50 | 50 | 214.0 | 0.00e+00 | 0.0000000 | **** | 2.9051852 | 8 , 30 | 5 | 7 | 3.0504444 | p < 1e-04 |
| unique_start_positions_coverage | 8 | 60 | 50 | 50 | 367.5 | 0.00e+00 | 0.0000000 | **** | 3.0088889 | 8 , 60 | 5 | 8 | 3.1593333 | p < 1e-04 |
| unique_start_positions_coverage | 15 | 30 | 50 | 50 | 453.5 | 0.00e+00 | 0.0000010 | **** | 3.1125926 | 15, 30 | 6 | 7 | 3.2682222 | p < 1e-04 |
| unique_start_positions_coverage | 15 | 60 | 50 | 50 | 783.0 | 1.00e-03 | 0.0280000 |
|
3.2162963 | 15, 60 | 6 | 8 | 3.3771111 | p = 0.028 |
| unique_start_positions_coverage | 30 | 60 | 50 | 50 | 1739.5 | 6.96e-04 | 0.0194880 |
|
3.3200000 | 30, 60 | 7 | 8 | 3.4860000 | p = 0.019488 |
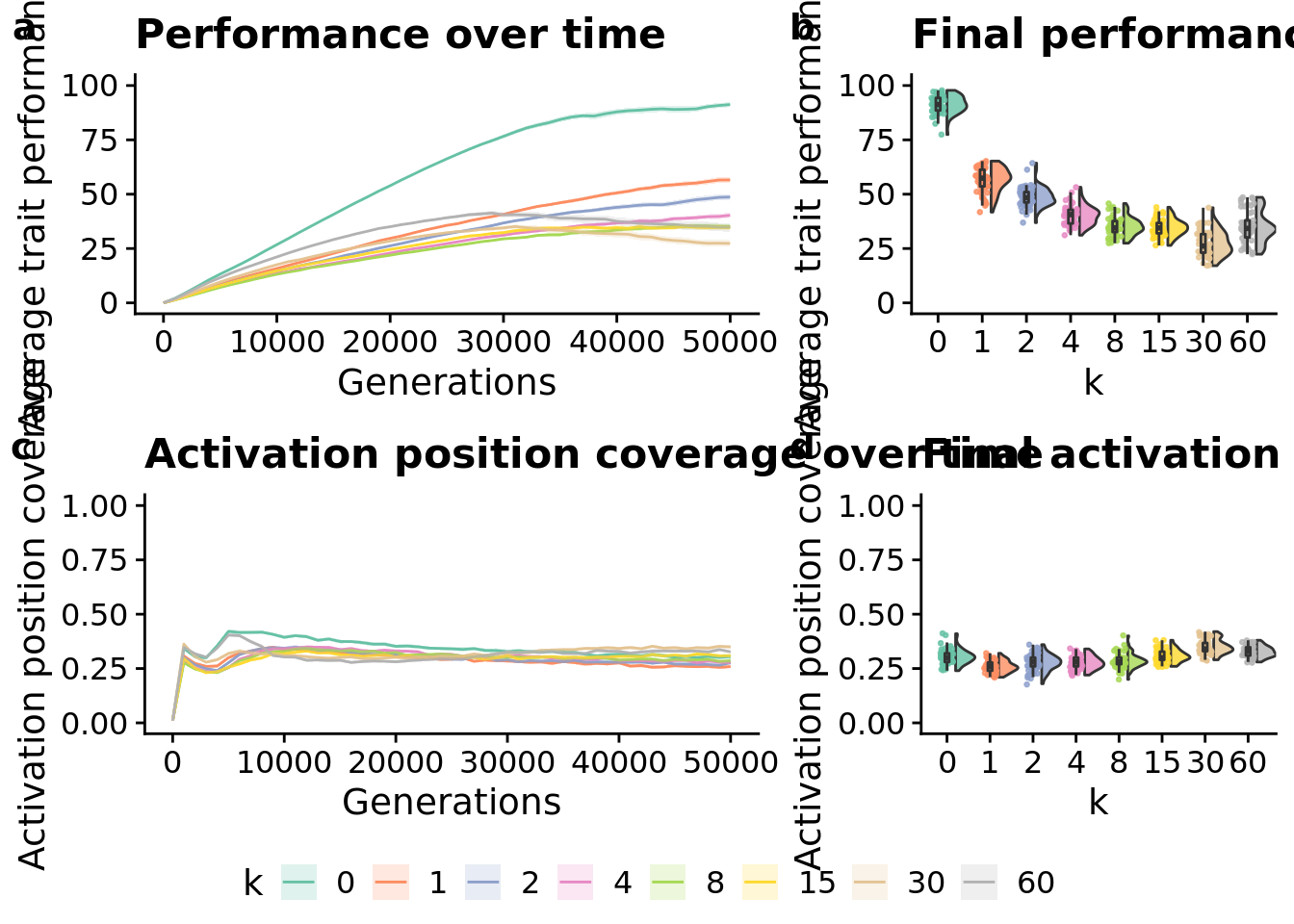
13.6 Manuscript figures
legend <- cowplot::get_legend(
elite_ave_performance_fig +
guides(
color=guide_legend(nrow=1),
fill=guide_legend(nrow=1)
) +
theme(
legend.position = "bottom",
legend.box="horizontal",
legend.justification="center"
)
)
grid <- plot_grid(
elite_ave_performance_fig +
ggtitle("Performance over time") +
theme(legend.position="none"),
elite_final_performance_fig +
ggtitle("Final performance") +
theme(),
unique_start_position_coverage_fig +
ggtitle("Activation position coverage over time") +
theme(legend.position="none"),
unique_start_positions_coverage_final_fig +
ggtitle("Final activation position coverage") +
theme(),
nrow=2,
ncol=2,
rel_widths=c(3,2),
labels="auto"
)
grid <- plot_grid(
grid,
legend,
nrow=2,
ncol=1,
rel_heights=c(1, 0.1)
)
save_plot(
paste(working_directory, "imgs/novelty-panel.pdf", sep=""),
grid,
base_width=12,
base_height=8
)
grid