Chapter 10 Cohort lexicase
10.1 Overview
10.2 Analysis dependencies
library(ggplot2)
library(tidyverse)
library(knitr)
library(cowplot)
library(viridis)
library(RColorBrewer)
library(rstatix)
library(ggsignif)
library(Hmisc)
library(kableExtra)
source("https://gist.githubusercontent.com/benmarwick/2a1bb0133ff568cbe28d/raw/fb53bd97121f7f9ce947837ef1a4c65a73bffb3f/geom_flat_violin.R")These analyses were conducted in the following computing environment:
## _
## platform x86_64-pc-linux-gnu
## arch x86_64
## os linux-gnu
## system x86_64, linux-gnu
## status
## major 4
## minor 1.0
## year 2021
## month 05
## day 18
## svn rev 80317
## language R
## version.string R version 4.1.0 (2021-05-18)
## nickname Camp Pontanezen10.3 Setup
data_loc <- paste0(working_directory, "data/timeseries.csv")
data <- read.csv(data_loc, na.strings="NONE")
data$cardinality <- as.factor(
data$OBJECTIVE_CNT
)
data$selection_name <- as.factor(
data$selection_name
)
data$epsilon <- as.factor(
data$LEX_EPS
)
data$proportion <- factor(
data$COH_LEX_PROP,
levels=c(1, 0.5, 0.2, 0.1, 0.05, 0.02, 0.01)
)
data$elite_trait_avg <-
data$ele_agg_per / data$OBJECTIVE_CNT
data$unique_start_positions_coverage <-
data$uni_str_pos / data$OBJECTIVE_CNT
final_data <- filter(data, evaluations==max(data$evaluations))
# Labeler for stats annotations
p_label <- function(p_value) {
threshold = 0.0001
if (p_value < threshold) {
return(paste0("p < ", threshold))
} else {
return(paste0("p = ", p_value))
}
}
# Significance threshold
alpha <- 0.05
####### misc #######
# Configure our default graphing theme
theme_set(theme_cowplot())10.4 Exploration diagnostic performance
elite_ave_performance_fig <- ggplot(
data,
aes(
x=evaluations,
y=elite_trait_avg,
color=proportion,
fill=proportion
)
) +
stat_summary(geom="line", fun=mean) +
stat_summary(
geom="ribbon",
fun.data="mean_cl_boot",
fun.args=list(conf.int=0.95),
alpha=0.2,
linetype=0
) +
scale_y_continuous(
name="Average trait performance",
limits=c(0, 100)
) +
scale_x_continuous(
name="Evaluations"
) +
scale_fill_brewer(
name="Cohort size",
palette=cb_palette
) +
scale_color_brewer(
name="Cohort size",
palette=cb_palette
)
elite_ave_performance_fig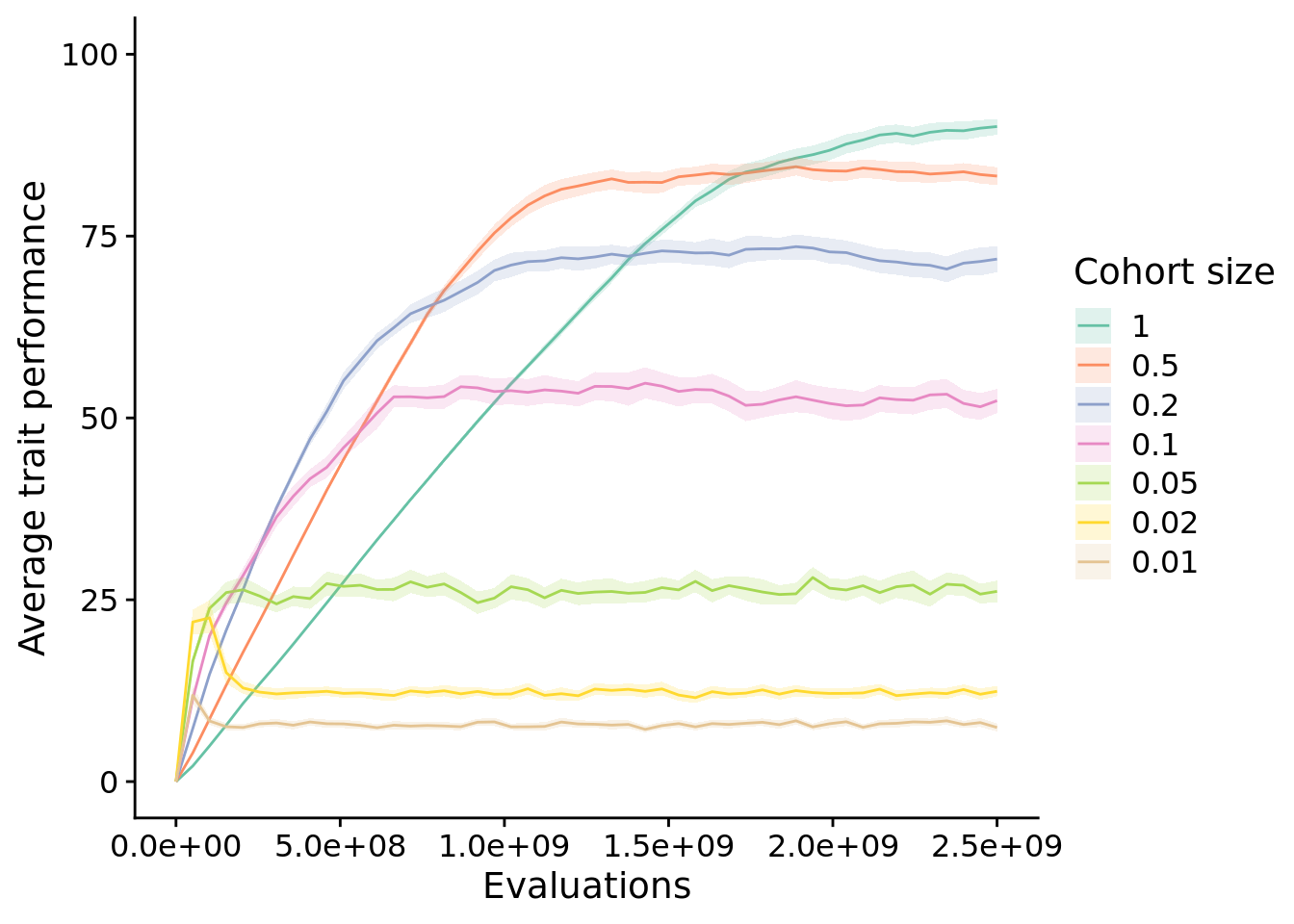
10.4.1 Final performance
# Compute manual labels for geom_signif
stat.test <- final_data %>%
wilcox_test(elite_trait_avg ~ proportion) %>%
adjust_pvalue(method = "bonferroni") %>%
add_significance() %>%
add_xy_position(x="proportion",step.increase=1)
stat.test$manual_position <- stat.test$y.position * 1.05
stat.test$label <- mapply(p_label,stat.test$p.adj)elite_final_performance_fig <- ggplot(
final_data,
aes(x=proportion, y=elite_trait_avg, fill=proportion)
) +
geom_flat_violin(
position = position_nudge(x = .2, y = 0),
alpha = .8,
scale="width"
) +
geom_point(
mapping=aes(color=proportion),
position = position_jitter(width = .15),
size = .5,
alpha = 0.8
) +
geom_boxplot(
width = .1,
outlier.shape = NA,
alpha = 0.5
) +
scale_y_continuous(
name="Average trait performance",
limits=c(0, 100)
) +
scale_x_discrete(
name="Cohort size"
) +
scale_fill_brewer(
name="Cohort size",
palette=cb_palette
) +
scale_color_brewer(
name="Cohort size",
palette=cb_palette
) +
theme(
legend.position="none"
)
elite_final_performance_fig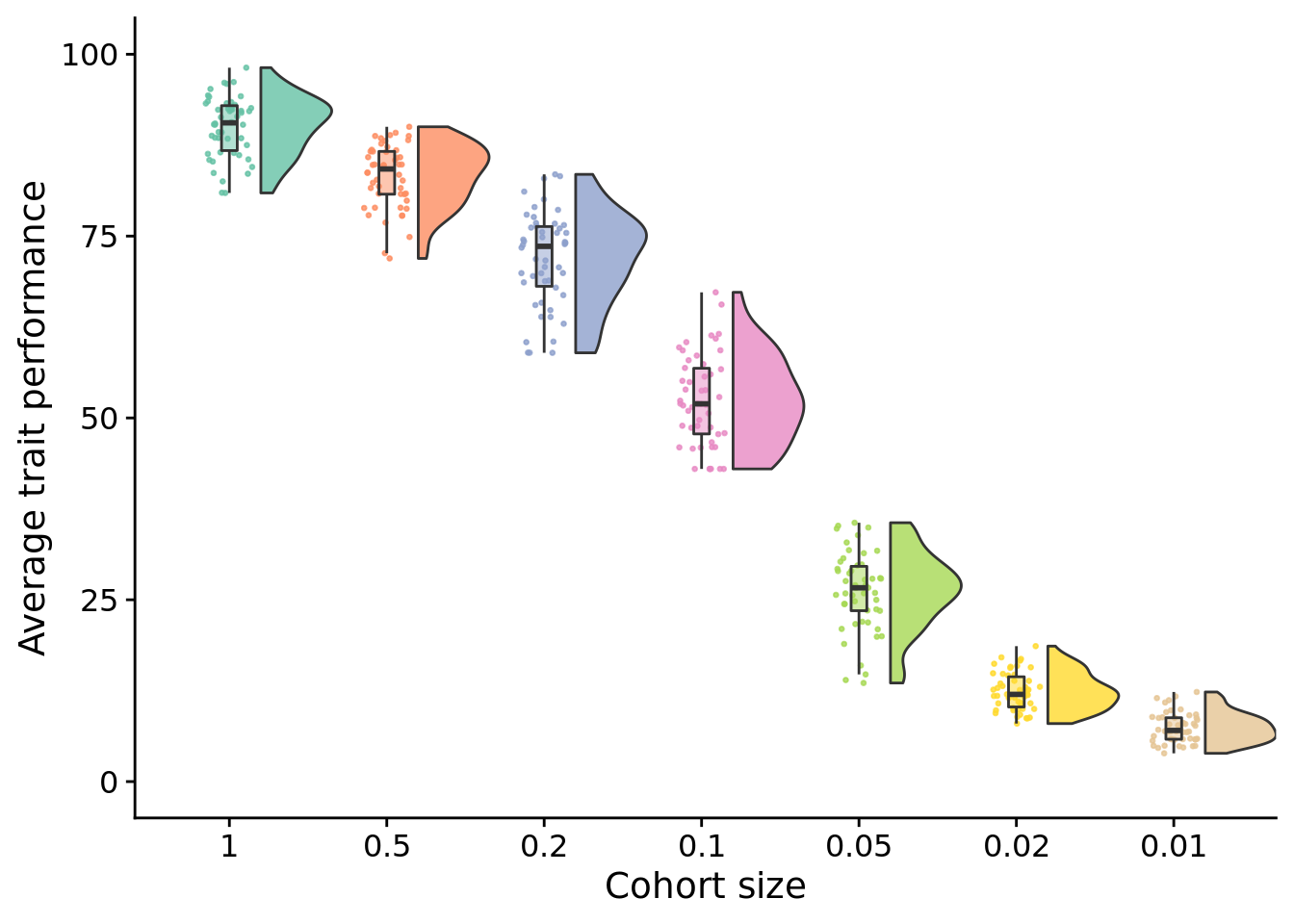
stat.test %>%
kbl() %>%
kable_styling(
bootstrap_options = c(
"striped",
"hover",
"condensed",
"responsive"
)
) %>%
scroll_box(width="600px")| .y. | group1 | group2 | n1 | n2 | statistic | p | p.adj | p.adj.signif | y.position | groups | xmin | xmax | manual_position | label |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| elite_trait_avg | 1 | 0.5 | 50 | 50 | 2173 | 0 | 0 | **** | 183.9910 | 1 , 0.5 | 1 | 2 | 193.1906 | p < 1e-04 |
| elite_trait_avg | 1 | 0.2 | 50 | 50 | 2489 | 0 | 0 | **** | 274.1178 | 1 , 0.2 | 1 | 3 | 287.8236 | p < 1e-04 |
| elite_trait_avg | 1 | 0.1 | 50 | 50 | 2500 | 0 | 0 | **** | 364.2445 | 1 , 0.1 | 1 | 4 | 382.4567 | p < 1e-04 |
| elite_trait_avg | 1 | 0.05 | 50 | 50 | 2500 | 0 | 0 | **** | 454.3712 | 1 , 0.05 | 1 | 5 | 477.0898 | p < 1e-04 |
| elite_trait_avg | 1 | 0.02 | 50 | 50 | 2500 | 0 | 0 | **** | 544.4980 | 1 , 0.02 | 1 | 6 | 571.7229 | p < 1e-04 |
| elite_trait_avg | 1 | 0.01 | 50 | 50 | 2500 | 0 | 0 | **** | 634.6247 | 1 , 0.01 | 1 | 7 | 666.3560 | p < 1e-04 |
| elite_trait_avg | 0.5 | 0.2 | 50 | 50 | 2333 | 0 | 0 | **** | 724.7515 | 0.5, 0.2 | 2 | 3 | 760.9891 | p < 1e-04 |
| elite_trait_avg | 0.5 | 0.1 | 50 | 50 | 2500 | 0 | 0 | **** | 814.8782 | 0.5, 0.1 | 2 | 4 | 855.6222 | p < 1e-04 |
| elite_trait_avg | 0.5 | 0.05 | 50 | 50 | 2500 | 0 | 0 | **** | 905.0050 | 0.5 , 0.05 | 2 | 5 | 950.2553 | p < 1e-04 |
| elite_trait_avg | 0.5 | 0.02 | 50 | 50 | 2500 | 0 | 0 | **** | 995.1318 | 0.5 , 0.02 | 2 | 6 | 1044.8883 | p < 1e-04 |
| elite_trait_avg | 0.5 | 0.01 | 50 | 50 | 2500 | 0 | 0 | **** | 1085.2585 | 0.5 , 0.01 | 2 | 7 | 1139.5214 | p < 1e-04 |
| elite_trait_avg | 0.2 | 0.1 | 50 | 50 | 2450 | 0 | 0 | **** | 1175.3853 | 0.2, 0.1 | 3 | 4 | 1234.1545 | p < 1e-04 |
| elite_trait_avg | 0.2 | 0.05 | 50 | 50 | 2500 | 0 | 0 | **** | 1265.5120 | 0.2 , 0.05 | 3 | 5 | 1328.7876 | p < 1e-04 |
| elite_trait_avg | 0.2 | 0.02 | 50 | 50 | 2500 | 0 | 0 | **** | 1355.6388 | 0.2 , 0.02 | 3 | 6 | 1423.4207 | p < 1e-04 |
| elite_trait_avg | 0.2 | 0.01 | 50 | 50 | 2500 | 0 | 0 | **** | 1445.7655 | 0.2 , 0.01 | 3 | 7 | 1518.0538 | p < 1e-04 |
| elite_trait_avg | 0.1 | 0.05 | 50 | 50 | 2500 | 0 | 0 | **** | 1535.8923 | 0.1 , 0.05 | 4 | 5 | 1612.6869 | p < 1e-04 |
| elite_trait_avg | 0.1 | 0.02 | 50 | 50 | 2500 | 0 | 0 | **** | 1626.0190 | 0.1 , 0.02 | 4 | 6 | 1707.3200 | p < 1e-04 |
| elite_trait_avg | 0.1 | 0.01 | 50 | 50 | 2500 | 0 | 0 | **** | 1716.1457 | 0.1 , 0.01 | 4 | 7 | 1801.9530 | p < 1e-04 |
| elite_trait_avg | 0.05 | 0.02 | 50 | 50 | 2456 | 0 | 0 | **** | 1806.2725 | 0.05, 0.02 | 5 | 6 | 1896.5861 | p < 1e-04 |
| elite_trait_avg | 0.05 | 0.01 | 50 | 50 | 2500 | 0 | 0 | **** | 1896.3992 | 0.05, 0.01 | 5 | 7 | 1991.2192 | p < 1e-04 |
| elite_trait_avg | 0.02 | 0.01 | 50 | 50 | 2343 | 0 | 0 | **** | 1986.5260 | 0.02, 0.01 | 6 | 7 | 2085.8523 | p < 1e-04 |
10.5 Activation position coverage
unique_start_position_coverage_fig <- ggplot(
data,
aes(
x=evaluations,
y=unique_start_positions_coverage,
color=proportion,
fill=proportion
)
) +
stat_summary(geom="line", fun=mean) +
stat_summary(
geom="ribbon",
fun.data="mean_cl_boot",
fun.args=list(conf.int=0.95),
alpha=0.2,
linetype=0
) +
scale_y_continuous(
name="Activation position coverage",
limits=c(0.0, 1.0)
) +
scale_x_continuous(
name="Evaluations"
) +
scale_fill_brewer(
name="Cohort size",
palette=cb_palette
) +
scale_color_brewer(
name="Cohort size",
palette=cb_palette
)
unique_start_position_coverage_fig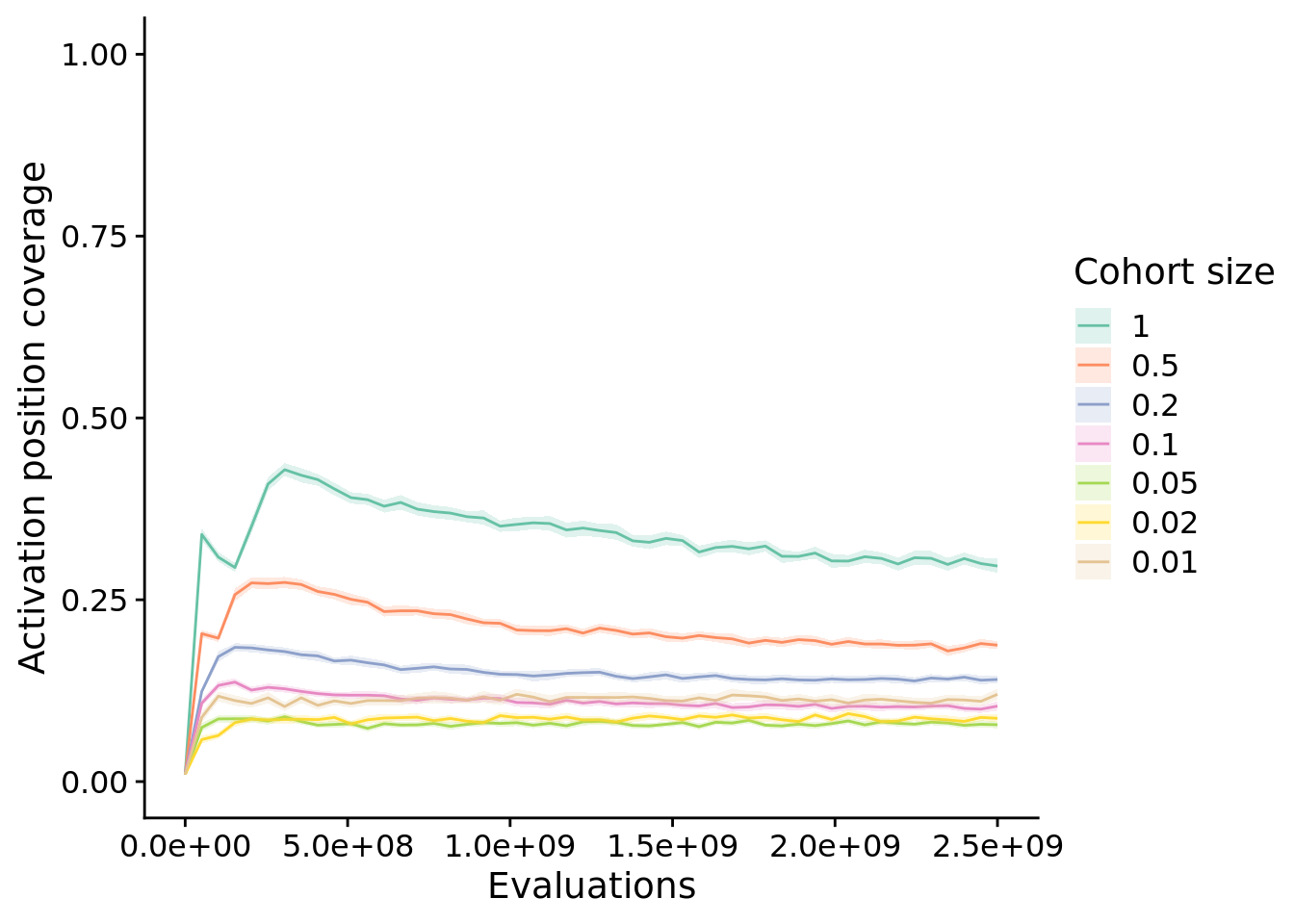
10.5.1 Final activation position coverage
# Compute manual labels for geom_signif
stat.test <- final_data %>%
wilcox_test(unique_start_positions_coverage ~ proportion) %>%
adjust_pvalue(method = "bonferroni") %>%
add_significance() %>%
add_xy_position(x="proportion",step.increase=1)
stat.test$manual_position <- stat.test$y.position * 1.05
stat.test$label <- mapply(p_label,stat.test$p.adj)unique_start_positions_coverage_final_fig <- ggplot(
final_data,
aes(
x=proportion,
y=unique_start_positions_coverage,
fill=proportion
)
) +
geom_flat_violin(
position = position_nudge(x = .2, y = 0),
alpha = .8,
scale="width"
) +
geom_point(
mapping=aes(color=proportion),
position = position_jitter(width = .15),
size = .5,
alpha = 0.8
) +
geom_boxplot(
width = .1,
outlier.shape = NA,
alpha = 0.5
) +
scale_y_continuous(
name="Activation position coverage",
limits=c(0, 1.0)
) +
scale_x_discrete(
name="Cohort size"
) +
scale_fill_brewer(
name="Cohort size",
palette=cb_palette
) +
scale_color_brewer(
name="Cohort size",
palette=cb_palette
) +
theme(
legend.position="none"
)
unique_start_positions_coverage_final_fig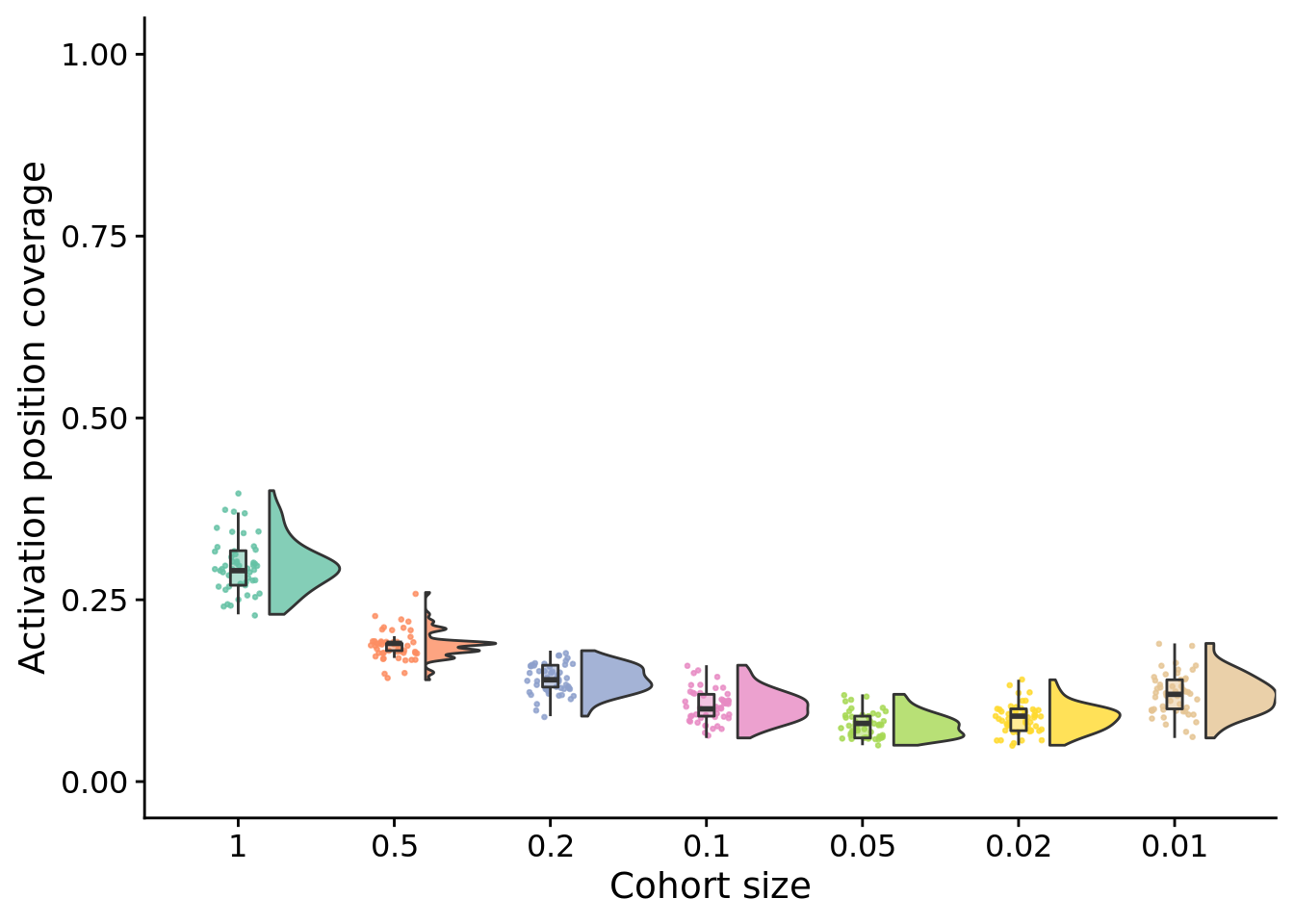
stat.test %>%
kbl() %>%
kable_styling(
bootstrap_options = c(
"striped",
"hover",
"condensed",
"responsive"
)
) %>%
scroll_box(width="600px")| .y. | group1 | group2 | n1 | n2 | statistic | p | p.adj | p.adj.signif | y.position | groups | xmin | xmax | manual_position | label |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| unique_start_positions_coverage | 1 | 0.5 | 50 | 50 | 2492.0 | 0.00e+00 | 0.0000000 | **** | 0.680 | 1 , 0.5 | 1 | 2 | 0.7140 | p < 1e-04 |
| unique_start_positions_coverage | 1 | 0.2 | 50 | 50 | 2500.0 | 0.00e+00 | 0.0000000 | **** | 0.974 | 1 , 0.2 | 1 | 3 | 1.0227 | p < 1e-04 |
| unique_start_positions_coverage | 1 | 0.1 | 50 | 50 | 2500.0 | 0.00e+00 | 0.0000000 | **** | 1.268 | 1 , 0.1 | 1 | 4 | 1.3314 | p < 1e-04 |
| unique_start_positions_coverage | 1 | 0.05 | 50 | 50 | 2500.0 | 0.00e+00 | 0.0000000 | **** | 1.562 | 1 , 0.05 | 1 | 5 | 1.6401 | p < 1e-04 |
| unique_start_positions_coverage | 1 | 0.02 | 50 | 50 | 2500.0 | 0.00e+00 | 0.0000000 | **** | 1.856 | 1 , 0.02 | 1 | 6 | 1.9488 | p < 1e-04 |
| unique_start_positions_coverage | 1 | 0.01 | 50 | 50 | 2500.0 | 0.00e+00 | 0.0000000 | **** | 2.150 | 1 , 0.01 | 1 | 7 | 2.2575 | p < 1e-04 |
| unique_start_positions_coverage | 0.5 | 0.2 | 50 | 50 | 2412.5 | 0.00e+00 | 0.0000000 | **** | 2.444 | 0.5, 0.2 | 2 | 3 | 2.5662 | p < 1e-04 |
| unique_start_positions_coverage | 0.5 | 0.1 | 50 | 50 | 2492.5 | 0.00e+00 | 0.0000000 | **** | 2.738 | 0.5, 0.1 | 2 | 4 | 2.8749 | p < 1e-04 |
| unique_start_positions_coverage | 0.5 | 0.05 | 50 | 50 | 2500.0 | 0.00e+00 | 0.0000000 | **** | 3.032 | 0.5 , 0.05 | 2 | 5 | 3.1836 | p < 1e-04 |
| unique_start_positions_coverage | 0.5 | 0.02 | 50 | 50 | 2499.5 | 0.00e+00 | 0.0000000 | **** | 3.326 | 0.5 , 0.02 | 2 | 6 | 3.4923 | p < 1e-04 |
| unique_start_positions_coverage | 0.5 | 0.01 | 50 | 50 | 2417.5 | 0.00e+00 | 0.0000000 | **** | 3.620 | 0.5 , 0.01 | 2 | 7 | 3.8010 | p < 1e-04 |
| unique_start_positions_coverage | 0.2 | 0.1 | 50 | 50 | 2216.5 | 0.00e+00 | 0.0000000 | **** | 3.914 | 0.2, 0.1 | 3 | 4 | 4.1097 | p < 1e-04 |
| unique_start_positions_coverage | 0.2 | 0.05 | 50 | 50 | 2469.0 | 0.00e+00 | 0.0000000 | **** | 4.208 | 0.2 , 0.05 | 3 | 5 | 4.4184 | p < 1e-04 |
| unique_start_positions_coverage | 0.2 | 0.02 | 50 | 50 | 2416.5 | 0.00e+00 | 0.0000000 | **** | 4.502 | 0.2 , 0.02 | 3 | 6 | 4.7271 | p < 1e-04 |
| unique_start_positions_coverage | 0.2 | 0.01 | 50 | 50 | 1834.0 | 5.05e-05 | 0.0010605 | ** | 4.796 | 0.2 , 0.01 | 3 | 7 | 5.0358 | p = 0.0010605 |
| unique_start_positions_coverage | 0.1 | 0.05 | 50 | 50 | 2044.0 | 0.00e+00 | 0.0000007 | **** | 5.090 | 0.1 , 0.05 | 4 | 5 | 5.3445 | p < 1e-04 |
| unique_start_positions_coverage | 0.1 | 0.02 | 50 | 50 | 1781.5 | 2.17e-04 | 0.0045570 | ** | 5.384 | 0.1 , 0.02 | 4 | 6 | 5.6532 | p = 0.004557 |
| unique_start_positions_coverage | 0.1 | 0.01 | 50 | 50 | 812.0 | 2.00e-03 | 0.0420000 |
|
5.678 | 0.1 , 0.01 | 4 | 7 | 5.9619 | p = 0.042 |
| unique_start_positions_coverage | 0.05 | 0.02 | 50 | 50 | 902.5 | 1.50e-02 | 0.3150000 | ns | 5.972 | 0.05, 0.02 | 5 | 6 | 6.2706 | p = 0.315 |
| unique_start_positions_coverage | 0.05 | 0.01 | 50 | 50 | 242.0 | 0.00e+00 | 0.0000000 | **** | 6.266 | 0.05, 0.01 | 5 | 7 | 6.5793 | p < 1e-04 |
| unique_start_positions_coverage | 0.02 | 0.01 | 50 | 50 | 405.5 | 0.00e+00 | 0.0000001 | **** | 6.560 | 0.02, 0.01 | 6 | 7 | 6.8880 | p < 1e-04 |
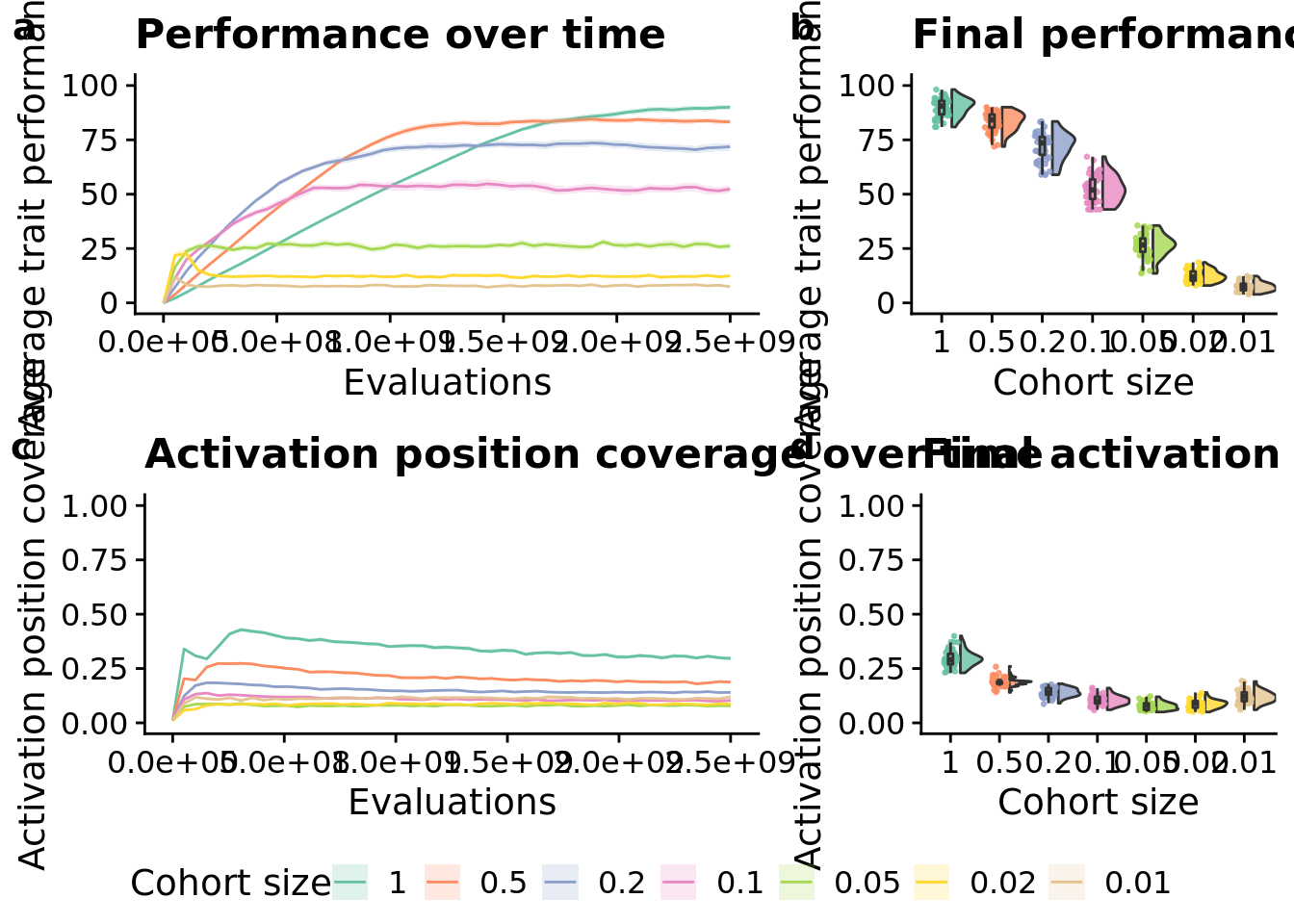
10.6 Manuscript figures
legend <- cowplot::get_legend(
elite_ave_performance_fig +
guides(
color=guide_legend(nrow=1),
fill=guide_legend(nrow=1)
) +
theme(
legend.position = "bottom",
legend.box="horizontal",
legend.justification="center"
)
)
grid <- plot_grid(
elite_ave_performance_fig +
ggtitle("Performance over time") +
theme(legend.position="none"),
elite_final_performance_fig +
ggtitle("Final performance") +
theme(),
unique_start_position_coverage_fig +
ggtitle("Activation position coverage over time") +
theme(legend.position="none"),
unique_start_positions_coverage_final_fig +
ggtitle("Final activation position coverage") +
theme(),
nrow=2,
ncol=2,
rel_widths=c(3,2),
labels="auto"
)
grid <- plot_grid(
grid,
legend,
nrow=2,
ncol=1,
rel_heights=c(1, 0.1)
)
save_plot(
paste(working_directory, "imgs/cohort-panel.pdf", sep=""),
grid,
base_width=12,
base_height=8
)
grid